Santa Cruz Mountains Trail Stewardship
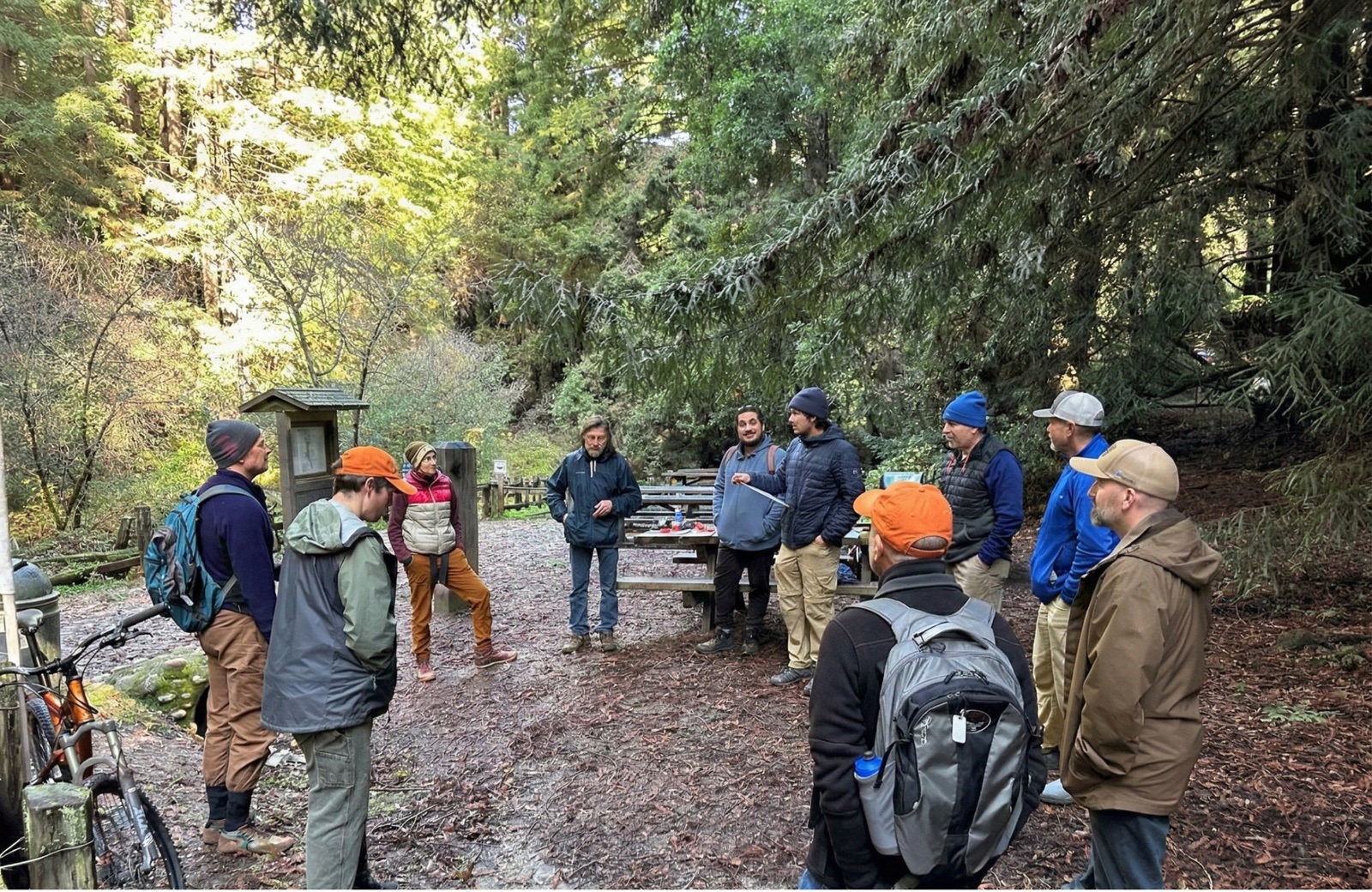
Part of SCMTS's volunteer trail crews building and maintaining trails at Wilder Ranch State Park and Henry Cowell Redwoods State Park — contributing to new public trail access in the Santa Cruz Mountains.
- Trail construction and maintenance on state park lands
- Contributing to expanded public access and conservation corridor connectivity
- Part of a 25+ year legacy of volunteer-led trail building in the region